News
The Wall Lab showcases latest research at the ICML 2023 Workshop
February 16, 2024
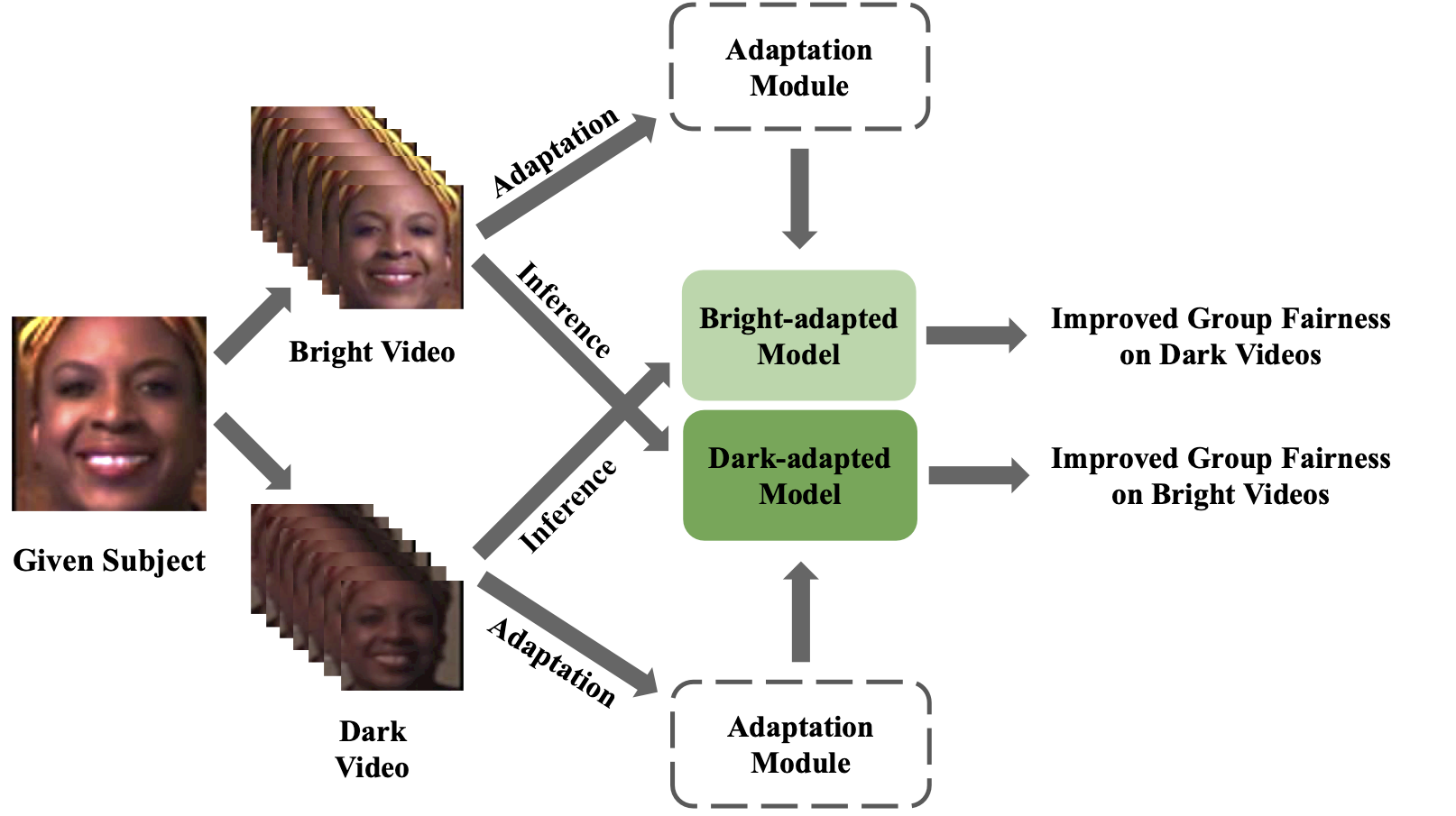
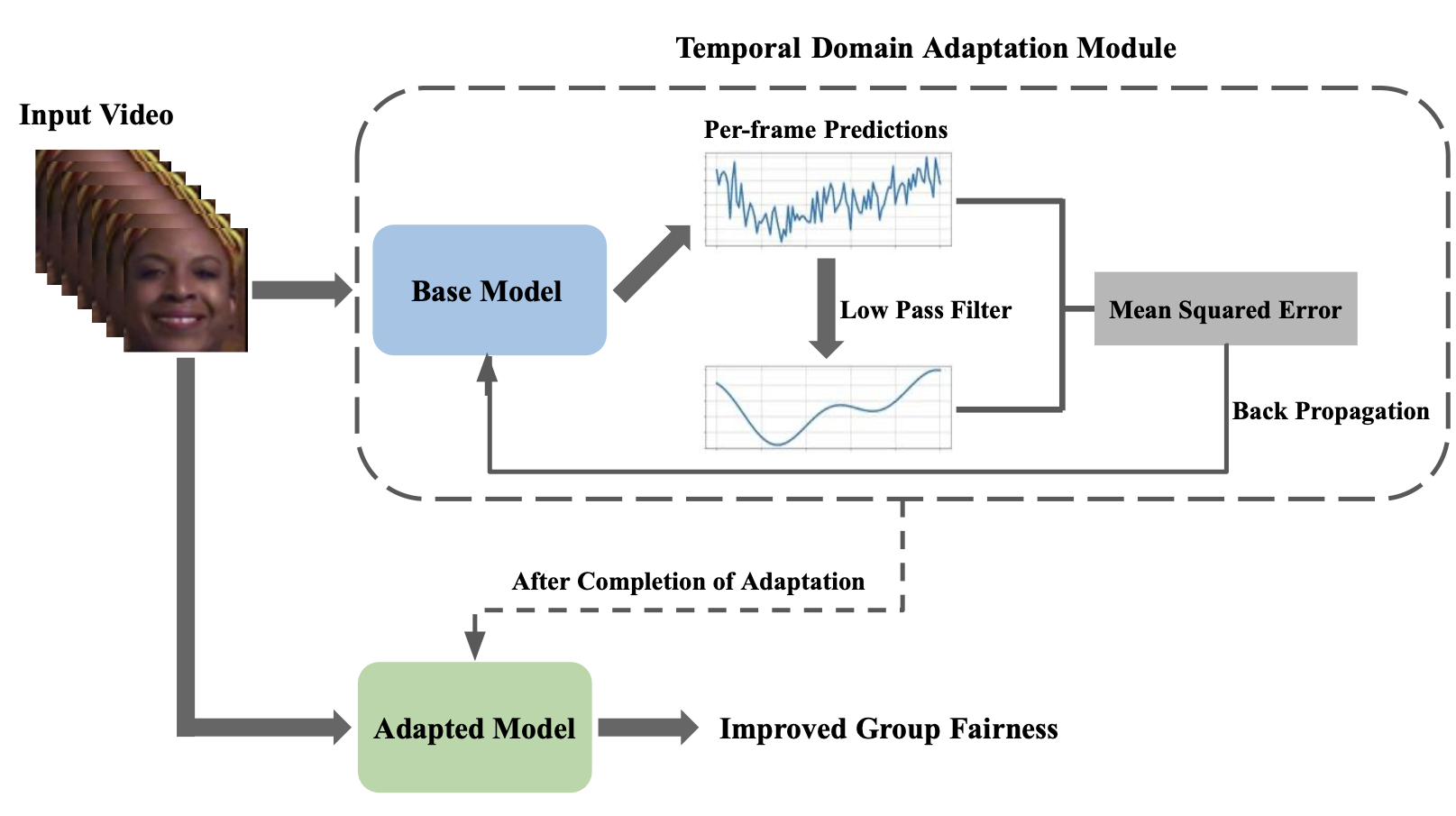
The Wall Lab at ICML 2023 Workshop on Spurious Correlations, Invariance, and Stability, showcased their latest research through a paper that introduces a novel unsupervised method aimed at enhancing fairness in AI models. By analyzing temporal coherence in video sequences, this approach promises to significantly improve predictive fairness across different demographic variables and lighting conditions, marking a meaningful step towards more equitable and personalized AI solutions.

Will you use Digital Health to Contribute to Autism Research?
November 19, 2019
In the last decade, autism spectrum disorder (ASD) prevalence in the U.S. increased by over 200%, affecting 1 in 40 children.1,2 Autism is estimated to affect 2% of children worldwide,2,3 and has become one of the most pressing pediatric health concerns globally.4,5,6,7 Standard approaches to diagnosis, such as the Autism Diagnostic Observation Schedule (ADOS)8 and the Autism Diagnostic Interview (ADI-R),9,10 and to therapy, such as Applied Behavioral Analysis (ABA), are difficult to access due to shortages of clinical practitioners, particularly in lower-income countries.9,11,12 The average age of diagnosis in the U.S. is 5 years of age for high-income and 8 years of age for low-income families. 2,13,14 Roughly 27% of U.S. children over the age of 8 remain undiagnosed.15
There is a high urgency to resolve these widespread problems with access to care. Research has shown that behavioral therapy by 5 years of age 17-21 can lessen or even eliminate core autism deficits including restricted and/or repetitive behaviors, difficulty with language, poor social attention, inability to understand facial expressions, and disinterest with social interactions.17,19,22-28 Worse, the impact of the interventions degrades after the age of 5, and by 8 years of age, children often do not respond as effectively. Therefore, early detection and therapy are both of vital importance.
Arguably, the most potent way to address this health crisis is via digital technologies. Children with ASD have shown high engagement with gamified systems which present an exciting opportunity for the development of novel diagnostic and therapeutic platform.29 As one example, researchers at King’s College London recently developed an interactive game, ECHOES, which allowed children to acquire social communication and emotional regulation skills through guided interactions with an intelligent virtual avatar.30 Multiple similar interventions have emerged over the past decade leveraging the latest technological developments in multitouch and interactive interfaces,31-34 humanoid-robot design,35 virtual and augmented reality systems.36-38



Virtually all interactive technological developments rely on extensive libraries of labeled images of human emotion. Nonetheless, children are significantly underrepresented in these sources and thus the classifiers trained on these databases are not optimized for pediatric autism research.39 Embracing the challenge, our team recently developed a Charades-style mobile game GuessWhat.stanford.edu, which engages children with autism and their families in a fun and convenient interaction that reinforces prosocial learning while simultaneously generating data for diagnostic and therapeutic AI development.39, 40 This is where you come in.
We need your help to develop the app’s capabilities! In an effort to corroborate and further increase GuessWhat?’s clinical efficacy, we are currently enrolling parents of children with Autism Spectrum Disorder (ASD) to participate in a fun research study!
Interested? Find out more by signing up on kidsfirst.stanford.edu/guesswhat!

Not a parent of a child with ASD? We need you too! Help us by simply playing the app (available for iOS and Android) and sharing with friends and family.
To stay tuned for updates, follow us on https://www.facebook.com/TheWallLab/
References
- Hertz-Picciotto I, Delwiche L. The Rise in Autism and the Role of Age at Diagnosis. Epidemiology 2009; 20(1): 84-90.
- Kogan MD, Vladutiu CJ, Schieve LA, et al. The Prevalence of Parent-Reported Autism Spectrum Disorder Among US Children. Pediatrics 2018; 142(6): e20174161.
- Hahler E-M, Elsabbagh M. Autism: A Global Perspective. Current Developmental Disorders Reports 2015; 2(1): 58-64.
- Dawson G. Why it’s important to continue universal autism screening while research fully examines its impact. JAMA pediatrics 2016; 170(6): 527-8.
- Piccininni C, Bisnaire L, Penner M. Cost-effectiveness of Wait Time Reduction for Intensive Behavioral Intervention Services in Ontario, Canada. JAMA Pediatrics 2017; 171(1): 23-30.
- Murray CJ, Lopez AD. Alternative projections of mortality and disability by cause 1990-2020: Global Burden of Disease Study. Lancet 1997; 349(9064): 1498-504.
- Whiteford HA, Degenhardt L, Rehm J, et al. Global burden of disease attributable to mental and substance use disorders: findings from the Global Burden of Disease Study 2010. The lancet 2013; 382(9904): 1575-86.
- Gotham K, Risi S, Pickles A, Lord C. The Autism Diagnostic Observation Schedule: revised algorithms for improved diagnostic validity. J Autism Dev Disord 2007; 37(4): 613-27.
- Lord C, Rutter M, Le Couteur A. Autism Diagnostic Interview-Revised: a revised version of a diagnostic interview for caregivers of individuals with possible pervasive developmental disorders. J Autism Dev Disord 1994; 24(5): 659-85.
- Poustka F, Lisch S, Ruhl D, Sacher A, Schmotzer G, Werner K. The standardized diagnosis of autism, Autism Diagnostic Interview-Revised: interrater reliability of the German form of the interview. Psychopathology 1996; 29(3): 145-53.
- Koegel LK, Koegel RL, Ashbaugh K, Bradshaw J. The importance of early identification and intervention for children with or at risk for autism spectrum disorders. International Journal of Speech-Language Pathology 2014; 16(1): 50-6.
- Lord C, Rutter M, DiLavore P, Risi S, Gotham K, Bishop S. Autism diagnostic observation schedule, (ADOS-2) modules 1-4. Los Angeles, California: Western Psychological Services 2012.
- Mazurek MO, Handen BL, Wodka EL, Nowinski L, Butter E, Engelhardt CR. Age at first autism spectrum disorder diagnosis: the role of birth cohort, demographic factors, and clinical features. J Dev Behav Pediatr 2014; 35(9): 561-9.
- Siklos S, Kerns KA. Assessing the diagnostic experiences of a small sample of parents of children with autism spectrum disorders. Res Dev Disabil 2007; 28(1): 9-22.
- Association AP. Diagnostic and statistical manual of mental disorders (DSM-5®). Arlington, VA: American Psychiatric Pub; 2013.
- Kamau LZ. Autism Spectrum Disorders (ASD) in Kenya: Barriers Encountered in Diagnosis, Treatment and Management. J Res Pharm Sci 2017; 3(7): 1-11.
- Dawson G. Early behavioral intervention, brain plasticity, and the prevention of autism spectrum disorder. Dev Psychopathol 2008; 20(3): 775-803.
- Dawson G, Jones EJH, Merkle K, et al. Early Behavioral Intervention Is Associated With Normalized Brain Activity in Young Children With Autism. Journal of the American Academy of Child and Adolescent Psychiatry 2012; 51(11): 1150-9.
- Dawson G, Rogers S, Munson J, et al. Randomized, controlled trial of an intervention for toddlers with autism: the Early Start Denver Model. Pediatrics 2010; 125(1): e17-23.
- Landa RJ. Efficacy of early interventions for infants and young children with, and at risk for, autism spectrum disorders. International Review of Psychiatry 2018; 30(1): 25-39.
- Phillips DA, Shonkoff JP. From neurons to neighborhoods: The science of early childhood development. Washington, D.C.: National Academies Press; 2000.
- Dawson G. Early behavioral intervention, brain plasticity, and the prevention of autism spectrum disorder. Development and psychopathology 2008; 20(03): 775-803.
- Dawson G, Webb SJ, McPartland J. Understanding the nature of face processing impairment in autism: insights from behavioral and electrophysiological studies. Dev Neuropsychol 2005; 27(3): 403-24.
- Howlin P, Goode S, Hutton J, Rutter M. Adult outcome for children with autism. J Child Psychol Psychiatry 2004; 45(2): 212-29.
- Landa RJ, Holman KC, Garrett-Mayer E. Social and communication development in toddlers with early and later diagnosis of autism spectrum disorders. Arch Gen Psychiatry 2007; 64(7): 853-64.
- Palumbo L, Burnett HG, Jellema T. Atypical emotional anticipation in high-functioning autism. Mol Autism 2015; 6(1): 47.
- Sasson NJ, Pinkham AE, Weittenhiller LP, Faso DJ, Simpson C. Context Effects on Facial Affect Recognition in Schizophrenia and Autism: Behavioral and Eye-Tracking Evidence. Schizophr Bull 2016; 42(3): 675-83.
- Xavier J, Vignaud V, Ruggiero R, Bodeau N, Cohen D, Chaby L. A Multidimensional Approach to the Study of Emotion Recognition in Autism Spectrum Disorders. Front Psychol 2015; 6: 1954.
- Kalantarian H, Washington P, Schwartz J, Daniels J, Haber N, Wall DP. Guess What? Towards Understanding Autism from Structured Video Using Facial Affect. J Healthcare Informatics Research 2019; 3: 43-66.
- Bernardini S, Porayska-Pomsta K, Smith TJ (2014) Echoes: an intelligent serious game for fostering social communication in children with autism. Inf Sci 264:41–60
- Battocchi A, Pianesi F, Tomasini D, Zancanaro M, Esposito G, Venuti P, Ben Sasson A, Gal E, Weiss PL (2009) Collaborative puzzle game: a tabletop interactive game for fostering collaboration in children with autism spectrum disorders (asd). In: Proceedings of the ACM International Conference on Interactive Tabletops and Surfaces, ACM, pp 197–204
- Feil-Seifer D, Matari´c MJ (2009) Toward socially assistive robotics for augmenting interventions for children with autism spectrum disorders. In: Experimental robotics, Springer, pp 201–210
- Gal E, Bauminger N, Goren-Bar D, Pianesi F, Stock O, Zancanaro M, Weiss PLT (2009) Enhancing social communication of children with high-functioning autism through a co-located interface. AI Soc 24(1):75
- Giusti L, Zancanaro M, Gal E, Weiss PLT (2011) Dimensions of collaboration on a tabletop interface for children with autism spectrum disorder. In: Proceedings of the SIGCHI Conference on Human Factors in Computing Systems, ACM, pp 3295–3304
- Scassellati B, Admoni H, Matari´c M (2012) Robots for use in autism research. Annu Rev Biomed Eng 14:275–294
- Daniels J, Schwartz J, Haber N, Voss C, Kline A, Fazel A,Washington P, De T, Feinstein C,Winograd T et al (2017) 5.13 Design and efficacy of a wearable device for social affective learning in children with autism. J Am Acad Child Adolesc Psychiatry 56(10):S257
- Voss C, Washington P, Haber N, Kline A, Daniels J, Fazel A, De T, McCarthy B, Feinstein C, Winograd T et al (2016) Superpower glass: delivering unobtrusive real-time social cues in wearable systems. In: Proceedings of the 2016 ACM International Joint Conference on Pervasive and Ubiquitous Computing: Adjunct, ACM, pp 1218–1226
- Washington P, Voss C, Kline A, Haber N, Daniels J, Fazel A, De T, Feinstein C, Winograd T, Wall D (2017) Superpowerglass: a wearable aid for the at-home therapy of children with autism. Proceedings of the ACM on Interactive, Mobile, Wearable and Ubiquitous Technologies 1(3):112
- Kalantarian H, Jedoui K, Washington P, Wall DP. A Mobile Game for Automatic Emotion-Labeling of Images. IEEE Transactions on Games 2018.
- Kalantarian H, Washington P, Schwartz J, Daniels J, Haber N, Wall DP. Guess What? Journal of Healthcare Informatics Research 2018.
Autism Glass in New York Times
July 17, 2019
The New York Times recently published an article on the Autism Glass project. Check out the article featuring our research below!
Google Glass May Have an Afterlife as a Device to Teach Autistic Children
Updates from the Past Few Months
July 10, 2019
The Wall Lab has been very busy these past few months with four new papers published:
First, “Outgroup Machine Learning Approach Identifies Single Nucleotide Variants in Noncoding DNA Associated with Autism Spectrum Disorder”, was published to the Pacific Symposium on Biocomputing. This paper investigates the role of noncoding variation in the ASD phenotype and shows the importance of the noncoding region and the utility of independent control groups in effectively linking genetic variation to disease phenotype for complex disorders through whole genome sequencing and machine learning models.
Next, “Detecting Developmental Delay and Autism Through Machine Learning Models Using Home Videos of Bangladeshi Children: Development and Validation Study” was published to the Journal of Medical Internet Research. This paper is a continuation of our previous work, in which we demonstrated the efficacy of machine learning classifiers to accelerate the process by collecting home videos of US-based children, identifying a reduced subset of behavioral features that are scored by untrained raters using a machine learning classifier to determine children’s “risk scores” for autism. Using videos of Bangladeshi children collected from Dhaka Shishu Children’s Hospital, we aim to scale our pipeline to another culture and other developmental delays, including speech and language conditions.
Another paper, entitled “Validity of Online Screening for Autism: Crowdsourcing Study Comparing Paid and Unpaid Diagnostic Tasks”, was published to the Journal of Medical Internet Research. This paper performed a series of studies to explore whether paid crowd workers on Amazon Mechanical Turk (AMT) and citizen crowd workers on a public website shared on social media can provide accurate online detection of autism, conducted via crowdsourced ratings of short home video clips.
And lastly, “Identification and Quantification of Gaps in Access to Autism Resources in the United States: An Infodemiological Study”, was published to the Journal of Medical Internet Research. Using the GapMap database, this paper quantifies the gaps in access to autism resources through computing the average distance between the nearest resource and individual with ASD as well as the relative disconnect between supply and demand of autism diagnostic resources across the U.S.
Check out these published papers below:
Outgroup Machine Learning Approach Identifies Single Nucleotide Variants in Noncoding DNA Associated with Autism Spectrum Disorder
Detecting Developmental Delay and Autism Through Machine Learning Models Using Home Videos of Bangladeshi Children: Development and Validation Study
Validity of Online Screening for Autism: Crowdsourcing Study Comparing Paid and Unpaid Diagnostic Tasks
Identification and Quantification of Gaps in Access to Autism Resources in the United States: An Infodemiological Study
Check back soon for more exciting updates!
The Future of Everything Radio Show - Featuring Dr. Dennis Wall
April 02, 2019
Happy World Autism Awareness Day!
The Wall Lab’s very own PI, Dr. Dennis Wall, was recently featured on the radio show, “The Future of Everything” with Dr. Russ Altman. There, they discuss about the bottlenecks surrounding autism diagnosis and the potential solutions that may alleviate these difficulties through the Wall Lab’s ongoing projects, including the mobile charades game called Guess What?
Check out the video recording/podcast of the radio show down below and don’t forget to share with your family and friends!
Facebook Video - Part 1
Facebook Video - Part 2
Soundcloud Podcast